The study of bacteria has entered a new technological era driven by advanced imaging methodologies. Modern high-resolution and label-free imaging systems now allow scientists to visualize bacterial cells at the nanoscale, monitor dynamic cellular processes in real time, and analyze metabolic responses without disrupting natural physiology. These innovations are reshaping microbiology research by providing unprecedented insights into bacterial structure, behavior, and adaptation mechanisms.
Introduction
Bacteria are highly dynamic and structurally complex microorganisms whose functional organization often occurs at nanometer scales. Traditional microscopy techniques, while foundational to microbiology, are limited by optical resolution constraints and often require invasive labeling procedures.
Recent developments in optical engineering, computational imaging, and biophysical instrumentation have revolutionized bacterial visualization. Super-resolution microscopy, quantitative phase imaging, Raman-based spectroscopy, and atomic force microscopy now enable researchers to study microbial systems with exceptional spatial precision and temporal resolution.
These technologies are not only improving structural analysis but also enabling real-time monitoring of bacterial metabolism, stress responses, biofilm formation, and antimicrobial interactions at the single-cell level.
Label-Free Imaging Technologies
Label-free imaging technologies are revolutionizing bacterial research by capturing native cellular states without fluorescent tags or invasive labels. These methods provide detailed insights into mass, morphology, metabolism, and mechanics, advancing diagnostics and antibiotic studies.
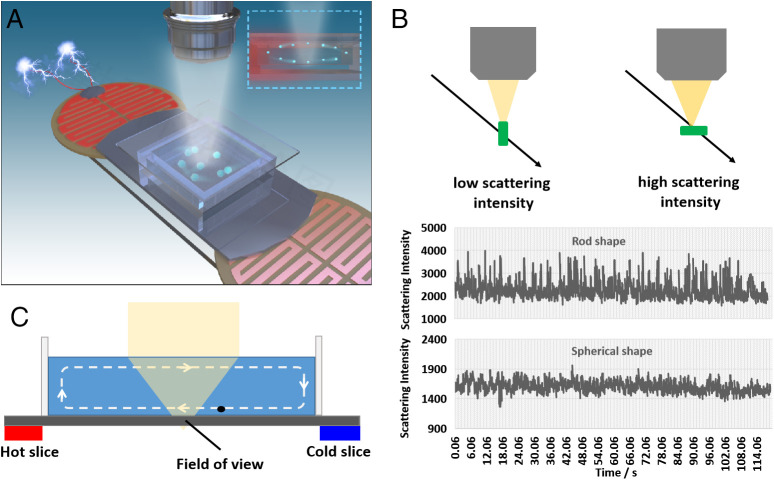
Figure:Bacteria detection principle by a single-particle imaging approach
Quantitative Phase Imaging (QPI)
QPI detects optical phase shifts from light passing through cells, generating maps of refractive index for dry mass and 3D morphology. It tracks real-time growth dynamics in single bacteria, such as antibiotic-induced slowdowns in E. coli within minutes. Holotomographic QPI variants enable pathogen identification via machine learning on 19 bacterial species.
Raman Spectroscopy and Coherent Raman
Raman spectroscopy reveals molecular "fingerprints" through inelastic light scattering, identifying biomolecules like proteins and lipids without labels. Coherent anti-Stokes Raman scattering (CARS) boosts speed and signal for live-cell metabolic profiling, distinguishing Staphylococcus from Pseudomonas by lipid content. Surface-enhanced Raman (SERS) substrates amplify signals for rapid species ID in clinical samples.
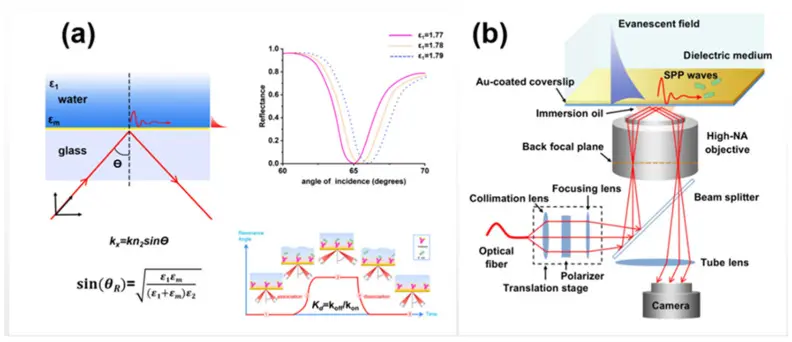
Figure:Schematic diagrams of (a) SPR optical system and (b) SPR microscopy.
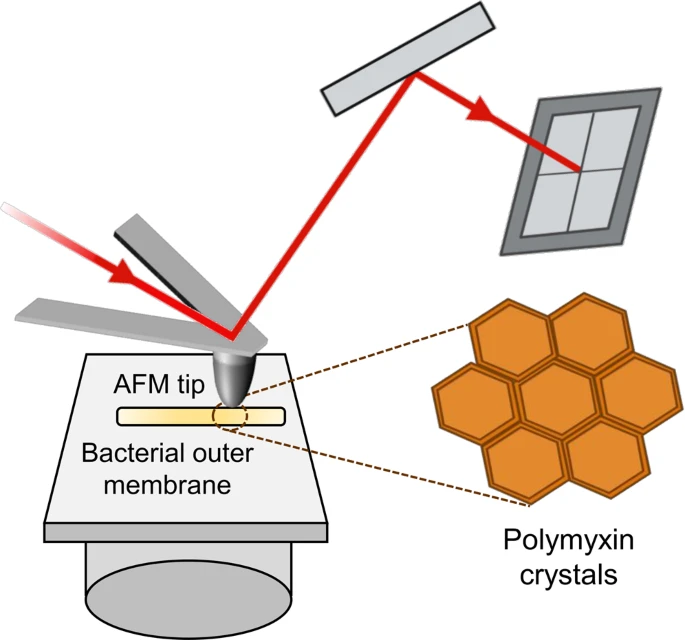
Atomic Force Microscopy (AFM)
AFM scans surfaces with a nanoscale probe to map topography, stiffness, and adhesion at angstrom resolution. It reveals bacterial envelope changes under antibiotics, like peptidoglycan softening in resistant strains. High-speed AFM captures dynamic processes, such as flagellar rotation or pore formation.
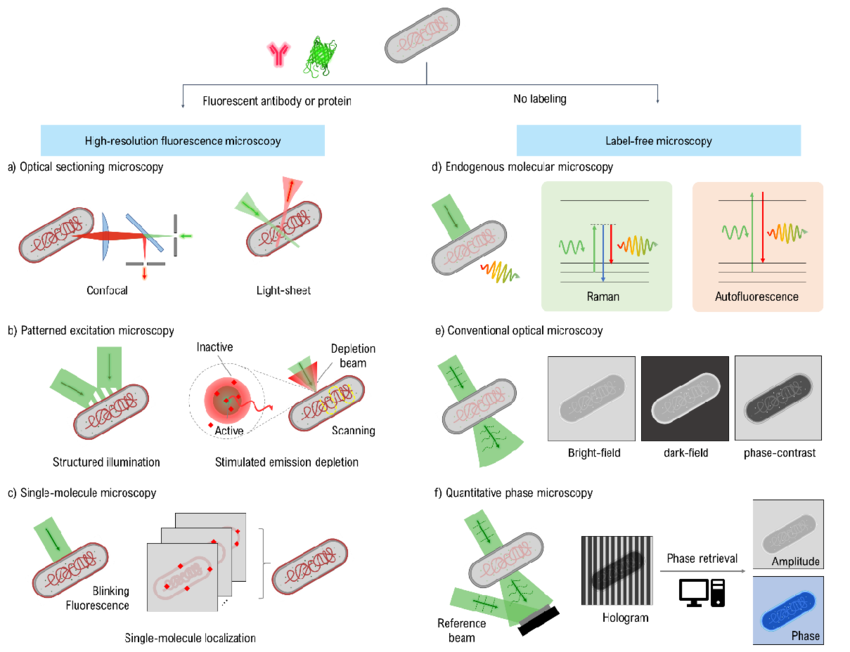
Figure: Schematic diagrams of high-resolution microscopy techniques are categorized based on their dependence on bacterial sample labeling.
Microfluidics and Single-Cell Platforms
Microfluidic devices paired with advanced microscopy offer unparalleled control over bacterial microenvironments, enabling long-term tracking of single cells. This integration reveals dynamic processes like antibiotic tolerance, phenotypic heterogeneity, and metabolic shifts without population averaging effects.
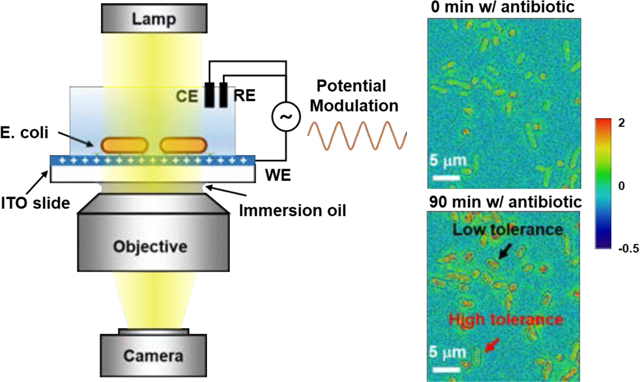
Figure: Single-Bacteria Impedance Mapping via Electro-Optical Impedance Microscopy
This figure illustrates the application of electro-optical impedance microscopy (EIM) for single-bacteria impedance imaging. Bacteria on an indium tin oxide–coated slide were subjected to low-frequency potential modulation while transmitted microscopy images were recorded. Fast Fourier transform analysis of the image sequences allowed separation of the DC component for cell morphology and the AC component for sub-cellular impedance mapping. The method reveals heterogeneous impedance responses of E. coli under different antibiotic treatments, particularly highlighting sensitivity to membrane-targeting antibiotics. EIM offers label-free, high-resolution imaging for monitoring bacterial viability and dynamic impedance changes at the single-cell level.
Core Advantages
Microfluidics deliver precise gradients of nutrients, antibiotics, or stressors at picoliter scales, while high-resolution imaging captures responses in real time. Trapping arrays immobilize individual bacteria for multi-hour observations, minimizing drift and phototoxicity. Time-lapse sequences quantify division rates, motility, and fluorescence from native autofluorescence or phase contrast.
Antibiotic Tolerance Studies
Mother-machine devices confine bacteria in growth channels, exposing them to antibiotics while imaging progeny. They demonstrate persister formation in E. coli, where subpopulations survive beta-lactams via slowed metabolism, reviving post-treatment. Dose-response curves reveal heterogeneity: some cells lyse rapidly, others enter dormancy, informing combination therapies.
Phenotypic Heterogeneity
Droplet microfluidics encapsulates single bacteria in oil-emulsion reactors, amplifying rare phenotypes like resistant mutants. Optical tweezers or dielectrophoresis sort cells by size/morphology during imaging, mapping noise in gene expression across isogenic populations. This uncovers bet-hedging strategies, where stochastic switching enhances survival under fluctuating conditions.
Stress Responses and Metabolism
Gradient generators simulate infection niches, tracking metabolic switching via Raman or QPI in Pseudomonas aeruginosa biofilms. Hypoxia chambers paired with multiphoton imaging show NADH/FAD autofluorescence shifts during anaerobiosis, linking redox states to virulence. Long-term platforms quantify lag phases and adaptive mutations under sublethal stress.
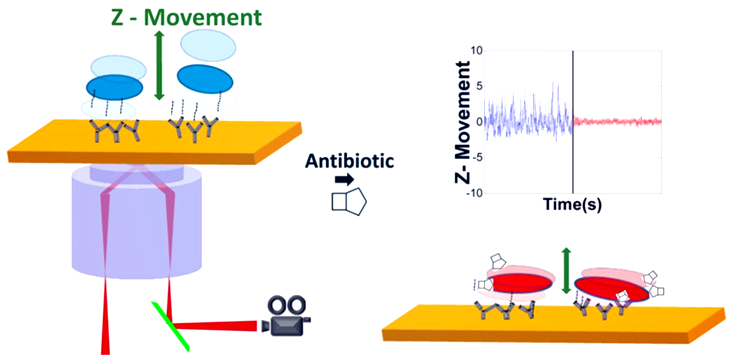
Figure: Schematic diagram of the rapid antimicrobial susceptibility test at single bacteria level using SPR microscopy
Technology Comparison
|
Platform |
Control Features |
Imaging Compatibility |
Key Insight Example |
|
Mother Machine |
Linear growth channels |
Phase/fluorescence |
Persister revival post-antibiotic |
|
Droplet Microfluidics |
Encapsulation, sorting |
Raman/dark-field |
Rare mutant enrichment |
|
Agarose Traps |
3D positioning |
Light-sheet |
Biofilm stress gradients |
|
Gradient Generators |
Chemical ramps |
QPI/STED |
Metabolic switching thresholds |
These platforms accelerate antibiotic susceptibility testing to under 1 hour and guide precision medicine against resistance. Hybrid systems with AI analysis promise scalable phenotyping for clinical isolates.
Impact on Microbiology Research
Advanced imaging technologies have dramatically transformed how microbiologists study bacteria, enabling deeper insights into structure, behavior, resistance mechanisms, and community dynamics at previously unattainable scales. Below are the major scientific contributions of these innovations, supported by recent research findings:
Enabling Nanoscale Structural Visualization
Modern imaging techniques such as super‑resolution microscopy, atomic force microscopy (AFM), and infrared nanospectroscopy allow scientists to visualize bacterial components at the nanoscale, surpassing traditional optical limits. These approaches reveal intricate details of cell walls, protein complexes, and antibiotic effects on membranes, providing new avenues for understanding bacterial architecture and drug interactions.
For example, Cryo‑electron microscopy (Cryo‑EM) enables reconstruction of bacterial ribosomes and antibiotic binding sites at near‑atomic resolution, deepening structural understanding.
Revealing Heterogeneity Within Bacterial Populations
Bacterial populations are often heterogeneous, meaning cells can differ significantly in behavior, metabolism, and drug response.
High‑resolution optical imaging methods help reveal this diversity by observing individual cells in real time. These technologies, especially when paired with computational analysis, enable researchers to monitor how different cells respond to stress, antibiotics, or environmental shifts insights that are crucial for precision microbiology.
Improving Analysis of Antimicrobial Resistance Mechanisms
Advanced imaging has been essential in decoding how bacteria adapt and resist antimicrobial agents.
Techniques like AFM and label‑free imaging have captured membrane changes and cell wall interactions under antibiotic exposure, shedding light on drug resistance pathways at molecular levels. These observations improve mechanistic understanding and inform targeted drug design.
Additionally, AI‑assisted imaging and pattern recognition enhance phenotypic antibiotic susceptibility testing by rapidly distinguishing resistant from sensitive strains.
Figure: In AFM imaging a sample is scanned with a sharp tip attached to a soft cantilever whose up- and down-deformations are detected by a laser-photodiode system.
Enhancing Biofilm Structural Studies
Biofilms protective communities of bacteria resist many conventional therapies. Advanced imaging tools like microfluidic microscopy and super‑resolution techniques now allow researchers to map biofilm architecture and dynamics in real time. This reveals how microorganisms communicate, adhere, and resist treatment, opening new strategies for disrupting persistent infections.
Review
Biofilms Exposed: Innovative Imaging and Therapeutic Platforms for Persistent Infections
by Manasi Haval et al.
Supporting Synthetic Biology and Microbial Engineering
In synthetic biology and microbial engineering, precise imaging is vital for monitoring engineered microbes, validating constructs, and optimizing designs.
High‑content and high‑resolution imaging provide quantitative data that guide modifications and ensure engineered strains behave as intended. When coupled with image‑based phenotyping, these tools can significantly speed up development cycles.
Computational Modeling and AI Integration
The combination of advanced imaging with artificial intelligence (AI) and machine learning (ML) has unlocked new capacities for high‑throughput analysis, automated feature extraction, and predictive modeling.
AI‑driven methods improve noise reduction, pattern recognition, and real‑time processing enabling researchers to analyze large imaging datasets at scale with greater accuracy and reproducibility.
AI models derived from imaging data are already being used to classify microbial phenotypes, detect antibiotic responses, and automate bacterial identification, illustrating how computational tools are reshaping microbial research workflows.
Conclusion
Advanced imaging techniques are redefining how bacteria are studied, shifting microbiology from population-based observation toward high-resolution, single-cell analysis. By integrating optical innovation, biophysical tools, and computational analytics, researchers can now observe bacterial systems with unprecedented clarity and precision.
These technological advances not only deepen fundamental scientific understanding but also support translational research in antimicrobial development, biotechnology, and environmental microbiology. As imaging systems continue to evolve, they will remain central to the next generation of discoveries in microbial science.