Welcome to the International Journal of Microbiology and Allied Sciences
IJOMAS is an international, peer-reviewed journal published quarterly, dedicated to sharing the latest discoveries in microbiology and related disciplines. Our goal is to provide an open, accessible platform for researchers, healthcare professionals, and students to share innovative research and foster global scientific collaboration.
Core Focus Areas
The journal accepts submissions across a wide range of microbiological fields:
Molecular Biology and Genetics
Microbial Physiology and Structural Biology
Microbial Biotechnology and Industrial Applications
Clinical, Food, Agricultural, and Environmental Microbiology
Microbial Ecology and Emerging Infectious Diseases
Bacteriology, Virology, Parasitology, Mycology
Immunology, Epidemiology, and Vaccinology

Mission and Vision
IJOMAS aims to accelerate the exchange of scientific knowledge by encouraging the publication of high-quality research and making information freely accessible to everyone. We see open science as a driver for innovation, academic dialogue, and global progress.

Open Access
All articles published in IJOMAS are under a Creative Commons (CC BY-NC 4.0) license, allowing readers to access, share, and adapt content while maintaining proper attribution. Each publication is archived in reliable digital repositories, ensuring long-term access and worldwide dissemination.

Who Can Contribute?
We welcome submissions from:
Researchers and academics
Healthcare professionals and clinicians
Graduate and postgraduate students
Interdisciplinary experts in microbiology and related fields
IJOMAS is more than a journal, it is a hub for discovery, innovation, and international scientific dialogue.
Advancing Microbiology Through Research
At IJOMAS, we bring together groundbreaking research from across the microbiology spectrum. Our mission is to connect scientists, clinicians, and students worldwide, making discoveries in microbiology and allied sciences accessible to everyone.
Key Research Areas
Welcome to the dynamic world of microbiology, where innovation meets discovery!
From the tiniest microbes to complex host interactions, the field is constantly evolving, reshaping medicine, biotechnology, and environmental science.
Antimicrobial Resistance (AMR) and Multidrug‑Resistant Infections
A major global scientific focus is understanding how microbes evolve resistance to antibiotics and developing new strategies to counteract this threat. Efforts include clinical perspectives on combating multidrug‑resistant bacteria and tracking resistance mechanisms at the molecular level
Core Mechanisms
Bacteria evolve resistance through enzymatic degradation (e.g., β-lactamases hydrolyzing antibiotics), target site alterations (e.g., modified penicillin-binding proteins in MRSA), reduced outer membrane permeability (e.g., porin loss in Gram-negatives), and efflux pumps (e.g., RND family expelling multiple drugs). Horizontal gene transfer via plasmids, transposons, and integrons rapidly disseminates these traits across species, amplified by environmental stressors like sub-lethal antibiotic exposure. Biofilms further enhance tolerance by creating physical barriers and metabolic dormancy, increasing resistance 10-1000-fold compared to planktonic cells
Table: Comprehensive table of antibiotics, their modes of action, and resistance mechanisms.
| Class | Mechanism of action | Target pathogens | Common resistance |
| β‐lactams | Inhibit cell wall synthesis via PBPs | Broad (Gram+ and Gram−) | β‐lactamases, altered PBPs, porin loss, efflux |
| Glycopeptides | Bind D‐Ala‐D‐Ala to block peptidoglycan synthesis | Gram+ (MRSA, VRE) | D‐Ala‐D‐Lac modification (VanA) |
| Macrolides | Inhibit 50S ribosome – block protein elongation | Gram+, atypicals (Mycoplasma, Chlamydia) | Efflux (msrA), ribosome methylation (erm) |
| Tetracyclines | Inhibit 30S ribosome – block tRNA binding | Broad‐spectrum incl. intracellular | Efflux (tetA), ribosome protection (tetM) |
| Aminoglycosides | Bind 30S ribosome – cause mRNA misreading | Aerobic Gram−, some Gram+ (Enterococcus) | Modifying enzymes (aac, ant), 16S methylation |
| Fluoroquinolones | Inhibit DNA gyrase & topoisomerase IV | Broad‐spectrum | gyrA/parC mutations, efflux (qnr), plasmids |
| Sulfonamides | Inhibit folic acid synthesis via DHPS | Broad, some protozoa | dhps mutations, PABA overproduction, efflux |
| Oxazolidinones | Inhibit 50S – block ribosome assembly | Gram+ (MRSA, VRE) | 23S rRNA mutation, cfr methylation |
| Polymyxins | Disrupt cell membrane (LPS interaction) | Gram− (Pseudomonas, Acinetobacter) | LPS modification (mcr‐1), efflux |
| Lincosamides | Inhibit 50S – block protein elongation | Gram+, anaerobes | erm‐mediated methylation, efflux |
| Rifamycins | Inhibit RNA polymerase – block transcription | TB, Neisseria, Staphylococcus | rpoB mutations, efflux |
| Fusidane | Inhibits EF‐G – halts protein elongation | Staphylococcus, Corynebacterium | fusA mutations, efflux (fusB) |
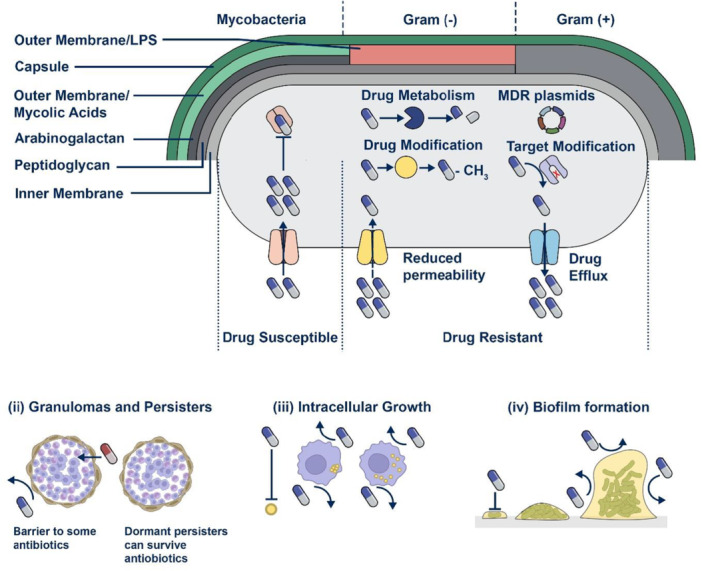
Figure:Structure of bacteria and classical antibiotic resistance mechanisms.
This image illustrates the structure of bacterial cells and the principal mechanisms underlying antibiotic resistance, showing how microorganisms survive antimicrobial exposure through efflux pumps that actively expel antibiotics, enzymatic degradation such as β-lactamase production that inactivates drugs, target modification including altered penicillin-binding proteins in MRSA, reduced membrane permeability in Gram-negative bacteria, and metabolic bypass pathways that circumvent inhibited biochemical processes. Together, these adaptive strategies contribute to the growing global antimicrobial resistance crisis and emphasize the urgent need for innovative therapeutic approaches and next-generation antimicrobial solutions.
Global Impact
AMR already contributes to nearly 5 million deaths yearly, straining healthcare with longer hospitalizations and higher costs, potentially reaching $100 trillion economically by 2050. MDR pathogens like carbapenem-resistant Enterobacteriaceae cause treatment failures over 50% in some regions, threatening surgeries, chemotherapy, and routine care. Low- and middle-income countries face heightened risks from poor surveillance, overuse in agriculture, and sanitation gaps.
Counterstrategies
Emerging approaches include phage therapy, antimicrobial peptides, quorum quenching to disrupt biofilms, and efflux pump inhibitors to restore antibiotic efficacy. One Health initiatives integrate human, animal, and environmental surveillance, while stewardship programs curb misuse; novel agents like nanobiotics and CRISPR-based editing target resistance genes directly. Only a few new antibiotic classes have emerged since 2010, underscoring needs for R&D incentives.
Machine Learning & AI in Microbiology
Artificial intelligence and deep learning are being applied to microbiological data in novel ways — from generating synthetic mass spectrometry spectra (MALDI‑TOF) to predicting microbial characteristics and pathogen risk based on metadata alone.
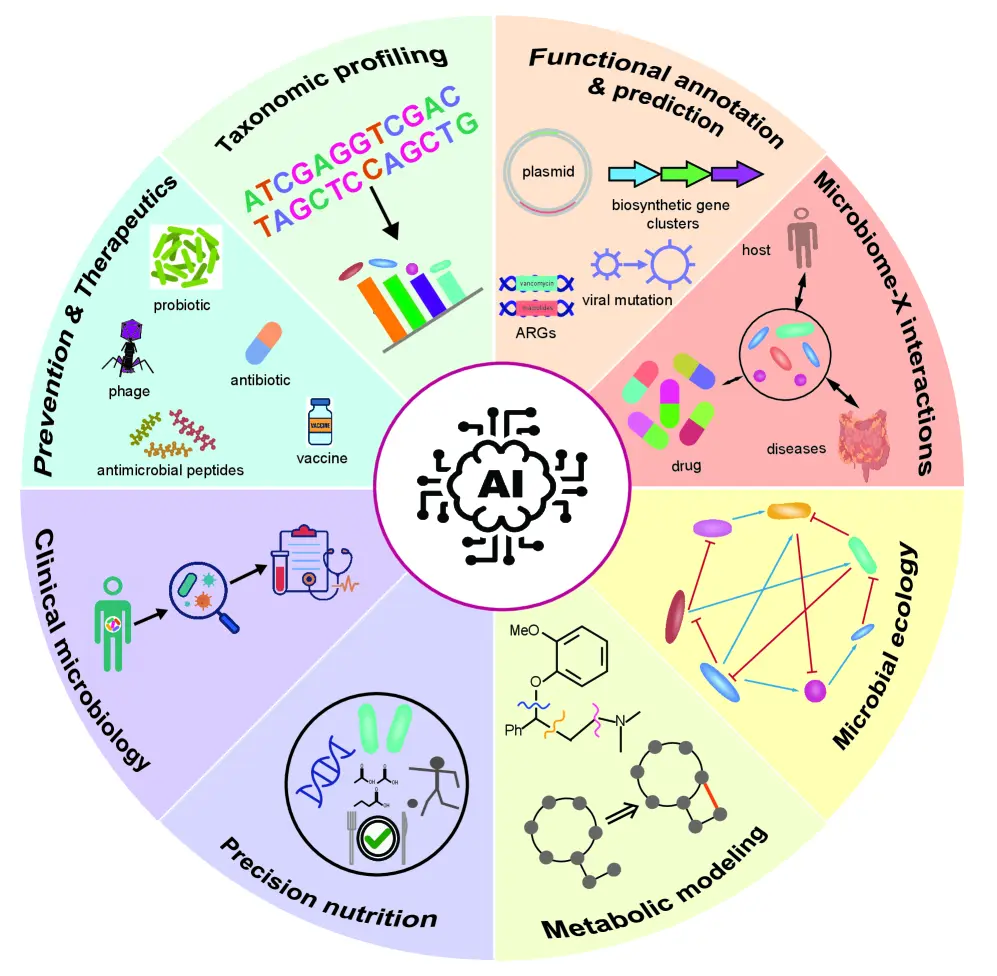
Figure:Application scenarios of AI in microbiology and microbiome research.
Artificial intelligence is revolutionizing microbiology and microbiome research by enabling advanced taxonomic profiling, functional gene and resistance prediction, and analysis of microbe–host and disease interactions. It strengthens microbial ecology and metabolic modeling, supports precision nutrition and personalized medicine, enhances clinical diagnostics and antimicrobial susceptibility testing, and accelerates innovation in therapeutics, including antibiotic discovery, probiotic development, phage therapy, and vaccine design.
MALDI-TOF Applications
Deep generative models like conditional VAEs (MALDIVAEs), GANs (MALDIGANs), and diffusion models (MALDIffusion) synthesize realistic MALDI-TOF mass spectra conditioned on microbial species labels. Trained on datasets like DRIAMS and MARISMa (>300,000 spectra), they reproduce peak structures and variability, enabling classifiers on synthetic data to match real-data performance (e.g., for E. coli, S. aureus). This augmentation mitigates class imbalance for rare pathogens, boosting accuracy up to 20% for underrepresented species like S. saprophyticus.
Metadata Predictions
Large language models (LLMs) classify microbial ontology (e.g., EMPO categories) and predict pathogen risks like E. coli contamination solely from environmental metadata (e.g., site, pH, temperature). LLMs outperform traditional models by reasoning over sparse data, generalizing across sites with AUC >0.85 for risk prediction. Applications span biosurveillance and ecology, integrating with AMR tracking via metadata-linked resistance forecasts.
Broader Impacts
AI excels in clinical microbiology tasks like organism ID, antimicrobial susceptibility, and outcome prediction from spectra or metagenomes. In AMR contexts, models detect resistance genes and predict susceptibility from MALDI-TOF peaks or sequences. Challenges include data imbalance and interpretability, addressed by hybrid models (e.g., CNN + RF).
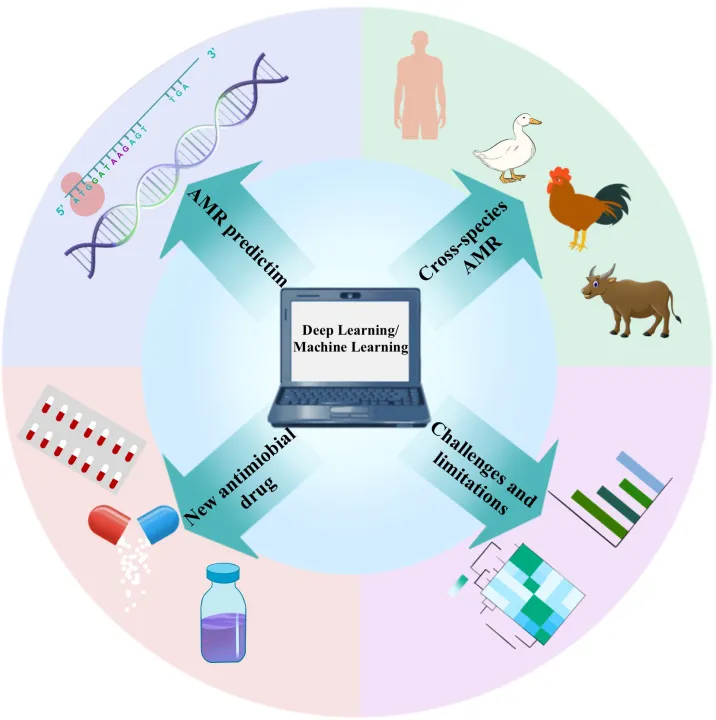
Figure:Deep Learning Strategies for Antimicrobial Resistance (AMR) Research.
Targeted deep learning models play a central role in antimicrobial resistance research. CNNs and RNNs (including LSTM and GRU) enable accurate AMR prediction from genetic sequences, while multi-task learning supports cross-species resistance analysis. In drug discovery, GANs and VAEs generate and optimize novel antimicrobial molecules. To improve transparency, explainable AI tools such as SHAP and LIME enhance the interpretability and reliability of complex predictive models
Conclusion
Microbiology stands at the intersection of innovation and global scientific progress. From tackling antimicrobial resistance to integrating artificial intelligence into microbial research, today’s discoveries are reshaping how we understand, analyze, and respond to the microbial world.
These emerging directions highlight a new era—where data science meets laboratory science, where molecular insights meet real-world applications, and where interdisciplinary collaboration drives meaningful breakthroughs.
At IJOMAS, we are committed to supporting and showcasing research that reflects this dynamic evolution. By embracing innovative methodologies and forward-thinking perspectives, we aim to contribute to a deeper scientific understanding of microorganisms and their impact across diverse ecosystems.
The future of microbiology is being written today — and it begins with discovery, innovation, and shared knowledge.